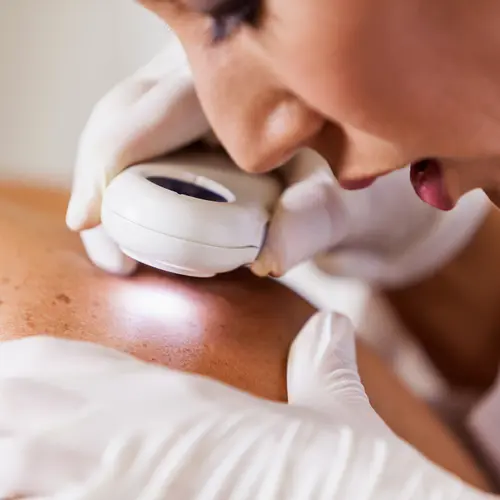
Skin cancer -- abnormal cell changes in the outer layer of skin -- is by far the most common cancer in the world. It can usually be cured, but the disease is a major health concern because it affects so many people. About half of fair-skinned people who live to age 65 will have at least one skin cancer. Most can be prevented by protecting your skin from the sun and ultraviolet rays.
Every malignant skin tumor will, over time, show up on the skin's surface. That makes this the only type of cancer that is almost always found in its early, curable stages.
Types of Skin Cancer
Skin cancers fall into two major categories: melanoma and nonmelanoma.
The most common skin cancers, basal cell carcinoma and squamous cell carcinoma, are nonmelanoma skin cancers and rarely life-threatening. They grow slowly, seldom spread beyond the skin, are easily found, and usually are cured. Basal cell carcinoma, which accounts for nearly 3 out of 4 skin cancers, is the slowest growing. Squamous cell carcinoma is somewhat more aggressive and more inclined to spread.
A rare nonmelanoma skin cancer is Kaposi's sarcoma, notable for its purple growths. It's related to a weak immune system and can be more serious. People with AIDS and the elderly tend to get it.
Some noncancerous skin growths could become cancerous. The most common are actinic keratoses -- crusty, reddish patches on sun-exposed skin that may scratch off but grow back.
The other type of skin cancer, melanoma, is a potentially aggressive, life-threatening cancer. It can start in dark skin tissue, such as a mole or birthmark, as well as in normally pigmented skin. For men, it generally shows up first on your head, neck, or between your shoulders and hips. Women tend to get it on their arms and legs. You may also find it on the palm of your hand, on the sole of your foot, under a fingernail or toenail, in mucus linings (in your mouth, vagina, or anus, for example), and even in your eye.
Melanoma is usually curable if found and treated early. But it grows faster than other types of skin cancer, and it can spread beyond your skin to other parts of the body, including your bones and brain. Then it's very hard to treat and is difficult to cure.
Causes
Spending too much time in the sun is the main cause of skin cancer. Sunlight has ultraviolet (UV) rays that can change the DNA in skin cells in ways that lead to cancer. Sunlamps, tanning booths, and X-rays also make these UV rays that damage skin.
Basal cell carcinoma and squamous cell carcinoma have been linked to ongoing sun exposure, typically in fair-skinned people who spend a lot of time outside. Melanoma has been linked with blistering sunburns; just one during childhood seems to double your risk for melanoma later in life.
Regularly working around certain chemicals and other things known to cause cancer may raise the odds of getting a nonmelanoma skin cancer, including:
- Coal tar
- Radium
- Insecticides with inorganic arsenic compounds
Who Gets Skin Cancer?
Skin cancer tends to affect people of light skin color because they're born with the least amount of protective melanin in their skin. The odds are highest if you're:
- Redheaded
- A blue-eyed blonde
- Someone with a pigment disorder, such as albinism
People with many freckles or moles, particularly odd-looking ones, may be vulnerable to melanoma. It's possible for dark-skinned people to get skin cancer, but it's rare and usually on lighter areas of their body, such as the soles of the feet or under fingernails or toenails.
Where you live also plays a role. Places with intense sunshine, such as Arizona and Hawaii, have a larger share of people with skin cancer. It's more common in places where fair-skinned people moved from less sunny areas, like Australia, which was settled largely by fair-skinned people of Irish and English descent.
About 3 times more men than women get skin cancer. It's more likely when you're older. Most people diagnosed are between ages 45 and 54, although more younger people are now being affected. If you or any close relatives have had skin cancer, your chances go up.
Treatment
Finding and treating skin cancer quickly is considered a cure. The type of skin cancer you have and how much it has spread, as well as your other health issues, will help your doctor decide how to treat it.
They may use one or more different ways to remove, kill, or stop the cancer cells from growing:
- Surgery
- Cryotherapy
- Radiation
- Chemotherapy or photochemotherapy
- Laser therapy
- Biologic or immunotherapy
- Targeted therapy
If the usual treatment doesn't work or it's difficult for you, you may be able to find a clinical trial. These test new ways to treat cancers that could be more effective or have lesser side effects.
Once you've had a skin cancer, like basal cell carcinoma, squamous cell carcinoma, and melanoma, and squamous cell carcinoma, there's a good chance that you'll develop a new skin cancer within a few years, so you'll need to get your skin checked regularly to catch it early.